« Previous 1 2 3 4 Next »
High-Performance Python – Distributed Python
UCX Python (UCX-Py)
The RAPIDS project develops Machine Learning tools and algorithms on top of GPUs, which is needed for high-speed networking for Dask (discussed in next section). Moreover, it needs a communication framework that can take advantage of GPU networking (NVLink). Because UCX provided these features, UCX Python, also referred to as UCX-Py, was created. Like RAPIDS and Dask, UCX-Py is an open source project under development. Akshay Venkatesh, one of the UCX-Py developers, presented a good introduction to UCX-Py at Nvidia GPU Technology Conference (GTC) 2019 (or you can view the slides online), where he presented some initial performance results comparing UCX to UCX-Py. For small messages, UCX-Py has only slightly more latency than UCX, but it is virtually identical to UCX for large messages. UCX-Py is just beginning. The first release, version 0.1, was announced on Septenber 28, 2019. You can install it with Conda,
$ conda create -n ucx -c conda-forge -c jakirkham/label/ucx cudatoolkit=ucx-proc=*=gpu ucx ucx-py python=3.7
or build it from source. UCX-Py has a serious role to play in allowing RAPIDS, Dask, and other tools to scale at the network level.
Dask
As data sets get constantly larger, at some point, they can easily become too large for a single node, resulting in the need to use multiple nodes just to hold and compute the data (parallel computing). The lead developer of Dask, Matthew Rocklin, gave a one-sentence overview of Dask: "Dask is a Python library for parallel programming that leverages task scheduling for computational problems." Dask’s initial focus was on machine learning, but it is now being applied to many areas of scientific or numerical computing. It is very Pythonic, allowing Python users to learn Dask quickly, and has been designed with three familiar interfaces: (1) array collections, (2) dataframe collections, and (3) bag collections.
One important aspect of Dask that often goes unnoticed is that the interfaces have been designed to work with "out-of-core" data – that is, data arrays that might be too large to keep in memory and "spill out" into data files. Dask arrays scale NumPy-like arrays or anything that looks like a NumPy array, such as the GPU-based CuPy, which takes NumPy code and makes it scalable across distributed systems. If you are familiar with NumPy coding, then using Dask arrays will not be a big change. This familiarity also extends to scikit-learn, which can use NumPy data types.
Dask dataframes, such as those from pandas or pandas-like structures (e.g., RAPIDS cuDF), scale, and you can use the pandas functions and methods with which you are already familiar. Dask bags are a bit different, in that they are not really designed to imitate a particular tool or data collection. Rather, they deal with collections of generic Python objects, occupying a small memory footprint and using Python iterators. You can think of it as a parallel version of PyToolz.
Dask bags allow you to use other data types or collections not covered by NumPy-like arrays or pandas-like dataframes. These high-level collections operate in parallel on datasets, even those that don't fit into main memory, and are very useful and important for large datasets in arrays, dataframes, or bags or when more performance is needed (i.e., running algorithms in parallel). As an example, you can use common pandas operations (e.g., groupby, join, and time series computations) in Dask dataframes. However, these are operated on in parallel for large data sets and for situations requiring higher performance.
Another example is for Dask bags, which are commonly used to parallelize simple computations on unstructured or semi-structured data, like text data, logfiles, JSON records, or user-defined Python objects. Operators such as map, filter, fold, and groupby can be used.
To better understand Dask, look at the simple example in Listing 1 that illustrates the close relationship between pandas and Dask.
Listing 1: Dask and pandas Similarities
| pandas | Dask |
import pandas as pd
df = pd.read_csv('2015-01-01.csv')
df.groupby(df.user_id).vaue.mean()
|
import dask.dataframe as dd
df = add.read_csv('2015-*-*.csv')
df.groupby(df.user_id).value.mean()
|
Porting code between the two is very easy. Dask is designed to run very fast and efficiently, especially on Dask arrays, Dask dataframes, and the following operations:
- Arithmetic
- Common aggregations
- Calls to apply
- Calls to value_counts() and drop_duplicates() or corr()
- Filtering with loc, isin, and row-wise selections
Beyond data collection, a second very important capability is the dynamic task scheduler that runs Python task graphs on distributed systems such as clusters. Dask allows custom, user-defined workloads, as well. During execution of the graph, Dask workers can be added or subtracted as needed, which is in contrast to other data messaging, in which the number of workers is, for the most part, fixed (e.g., with MPI, although it does have dynamic aspects). Losing an MPI rank during execution can be disastrous, whereas Dask can just continue using another worker and continue computing.
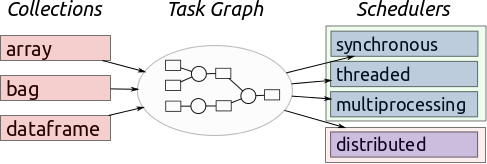
The schedulers can be broken into a few general categories: (1) synchronous, (2) threaded, (3) multiprocessing, and (4) distributed. These schedulers are designed to run on everything from a laptop for personal use, to thousand-node clusters. To run, Dask takes large-scale data collections and creates a task graph (Figure 1).
Each node of the task graph is a normal Python function, and the graph edges between nodes are normal Python objects created by one task as outputs and used by another task as inputs. Once these graphs are created, they need to be executed on parallel hardware. The task scheduler takes care of execution with of one of the previously mentioned schedulers.
The task scheduler(s) and execution graphs constitute a powerful workflow. For example, if a worker suddenly stops, the scheduler can just start using another worker, and no processing is lost. Also, if needed, additional workers can be used either for performance reasons or to provide more computational power if the graph needs it at some point.
Machines like laptops use a "single machine scheduler" that provides basic features on a local process or thread pool. It is simple and cheap to use for a single node because it doesn't have to scale. Don't forget that Dask can use out-of-core datasets, so you don't have to fit everything into memory. The performance may be a bit slower, but you can still run the workflows on your laptop.
Distributed resources use distributed schedulers, which are more sophisticated and require a requisite amount of effort to configure. The resulting workflow can then be run across distributed resources such as a cluster. To repeat: One of the really cool things about Dask is that it can expand to use additional resources (nodes) as needed to process the graph, or it can stop using resources if they aren't needed. This is a bit different from classic HPC distributed systems that usually use a fixed number of resources.
Dynamic resource usage doesn't usually fit into the HPC world because the sharing of resources can cause problems, such as the overuse of memory or the use of too many compute resources (cores); however, dask-mpi lets you use Dask in the HPC world from within an existing MPI environment. Dask takes some work and practice to learn. An online tutorial walks you through a simple Dask example with various options and talks about the effect on overall performance. A large number of tutorials and examples can be found around the web, so take a look at them to get started. The next few sections discuss the integration of Dask and … well, everything!
Dask and UCX
The recent UCX-Py (previously discussed) integration with Dask allows Python code executed via Dask to utilize UCX. If you have a fast network (e.g., InfiniBand), the Dask tasks that require coordination and communication are minimized as much as possible. A couple of fairly recent articles on high-performance networking with UCX and DGX and Python and GPUs briefly discuss the integration of UCX, via UCX-Py, and Dask.
Dask + RAPIDS
Pandas, scikit-learn, and NumPy were not designed to scale; they were designed for single-image systems (a single node). Dask, along with other tools, was designed to scale to multiple machines. Dask integrated with UCX results in a scalable set of open source Python-based tools for better performance and the use of large datasets. The obvious next step was to integrate all of this with RAPIDS, an open source set of tools for running Machine Learning (ML) applications and general numerical Python applications on GPUs. RAPIDS focuses on GPU-compatible versions of pandas, scikit-learn, and NumPy (the GPU version called CuPy).
« Previous 1 2 3 4 Next »