Scalability abhors serial computation, but parallel I/O can defeat those limitations.
Parallel I/O Chases Amdahl Away
Amdahl’s Law explains why the serial portion of applications don’t scale in performance as computing resources are added for the same problem size. The serial portions of an application can come from any number of issues. For example, network speeds are limited by the speed of light, so data communication between processes always experience some delay and is not affected by the number of processes. Moreover, processes might need to perform computations before either sending or receiving data from other processes. These computations, while scaling with the number of processes, has a lower limit that contributes to serial time.
One of the common sources of serial time is I/O. Applications have to read and write data independent of the number of processes. Most commonly in the case of a distributed application, I/O is done serially by one process. In this article, I discuss some ideas for going beyond serial I/O to parallel I/O with some scalability. The number of computational resources can determine how much I/O contributes to serial time and allow the application to continue scaling as the number of resources grow.
Before jumping in with both feet, I’ll define parallel I/O as different processes in a distributed or threaded application that perform I/O, primarily read and write, at the same time. The processes can be on one node or across multiple nodes, but each process or thread should have its own processing unit. The number of processes performing I/O do not have to equal the number of computational resources (i.e. Nproc != Mio where Nproc is the number of computational processes and Mio is the number of processes doing I/O.
Parallel I/O can apply to writing data to multiple files at the same time or to a single file, with no restriction on that number of files. However, be very cautious about parallel I/O to the same file (more on that in the next section).
Strictly speaking, if you perform I/O to a single file where I/O is serial, this isn’t parallel I/O. I hope to give you some ideas in this article about how to reduce I/O by performing parallel I/O. Although not possible with serial applications, it is possible for multiprocess applications.
Single-Process Application
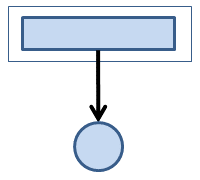
Many applications use just a single process (i.e., they are serial). The process performs any needed I/O (Figure 1). The box represents the system, and the file is represented by the circle (e.g., a hard drive).
I don’t use low-level libraries for performing I/O, but the general rule of thumb is to try not having more than one process write to the same file at the same time. Although people have told me they can do it, you must be amazingly cautious, and you have to understand filesystems at a high level. My gentle advice is, don’t do it.
Parallel Applications and Parallel I/O
Parallel applications use more than one process for computation. They could be processes or threads, but in this article, I will generically use the term “process.” The processes can be on a single node with one process using one core or thread, or with processes using cores on other nodes. I assume that each process has its own memory space. (I’m ignoring the case of shared memory for simplification.) To make life easier, I will also assume that the only way the processes share data is by communicating between themselves (e.g., Message Passing Interface (MPI)).
Performing I/O in a logical and coherent manner from disparate processes is not easy. It’s even more difficult to perform I/O in parallel. I’ll begin with a very simple parallel I/O paradigm.
File Per Process and Subsets of Processes
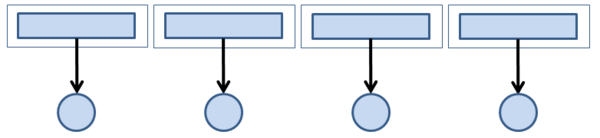
Probably the easiest way to perform parallel I/O is to have each process write or read to or from its own file. More precisely, an application with Nproc processes writes to Nfiles, where Nproc = Nfiles.
.
This scenario is very simple to code because each process does I/O to its own file, as in serial code, so you don’t need to change the I/O portion of the code. Moreover, there is no overlap, and all the I/O from Nproc processes is happening at the same time (in parallel).
The difficulty with this technique is that if you rerun the application, you must use the same number of processes as you used previously. If you ran with eight processes before, you have to run with eight processes again. For some users and applications this limitation isn’t an issue, but that means you can’t scale your application by using more processes. To do so, you would have to (1) somehow redistribute the data across Nnew processes (Nnew > Nprev), where Nprev is the previous number of processes or (2) rewrite the code to read the data from fewer files than processes (Mfiles < Nnew). In the next two subsections I examine those options.
Data Redistribution into Files
Having one process write and read to its own file is a great way for overall I/O performance to better scale with the number of processing elements. However, starting a run with Nproc processes but with only Mfiles files (Mfiles != Nproc) presents some difficulties. One way to solve this problem is to write an application that reads the data in Mfiles files and writes it into Nproc files.
A simple approach is to write an application outside of the computational application to perform the data redistribution before running another application that accesses the data. An advantage of this approach is that the computational code does not have to be rewritten. Most changes to the file format are handled in the data redistribution application, with perhaps a few small changes to the computational application when additional data is either read or written.
A good starting point for an external application is to write a serial application (one process) that reads the Mfiles data files into memory and then logically divides it into Nproc partitions and writes each one to a different file.
To illustrate this, two simple Python scripts are presented, in which one (Listing 1) writes random data to five files and the other (Listing 2) reads the data in the files and writes it to 11 files. The code in Listing 1 creates the five files that serve as input. A NumPy array of size nx by ny is created with random data and written to a unique file. This operation is repeated five times, creating five files that simulate a large 2D array of size nx by ny(nfile), where nfile is the number of files (in this case, five). Running this code should produce five files named file_Y.npy, where Y varies from 1 to 5.
Listing 1: Write to Five Files
'''
Create a numpy array of [nx x ny] random data and write it
to a file. Loop over this to create 5 files (for the sake
of this example)
'''
import numpy as np
# Define array size (nx x ny) per file
nx = 200
ny = 200
nfiles = 5 # Number of files to write
# Loop over number of files and write to files
for i in range(nfiles):
filename = "file_" + str(i) # filename
a = np.random.rand(nx,ny)*100.0 # Random data in arrayy
np.save(filename, a) # Write data to file
print(" Just finished writing file, ",filename,".npy")
# end forThe code in Listing 2 reads the five files, concatenates them to produce one large array named input_data, then partitions the array into nfiles_new partitions, with each partition written to a unique file. Note that the partitions might not be the same size because the data can’t be spread evenly over the 11 files.
Listing 2: Five Files In and 11 Files Out
'''
Read the data in "nfiles", subdivide into "nfiles_new"
paritions and write "nfiles_new" new files.
The array "input_data" holds all of the data from
the files in one array (ny x nx*nfiles). It is
paritioned and each partition is written to a unique
file.
'''
import numpy as np
import math
nfiles = 5 # Number of input files
# Read first file to start array
filename = "file_0.npy"
input_data = np.load(filename)
# Loop over remaining files and read the data
for i in range(1,nfiles):
# input filename
filename = "file_" + str(i) + ".npy"
a = np.load(filename) # Load the data into a
# Append a to input_data array
input_data = np.append(input_data, a, axis=1)
# end for
# Write to new files
nfiles_new = 11 # Number of new files to write
total_len = input_data.shape[1] # ny
# Number of new elements per file
n1_new = math.floor( total_len/nfiles_new)
print("n1_new = ",n1_new)
# Write to the first (nfiles_new-1) files in case the amount of
# data per file is not even
istart = 0
for j in range(nfiles_new-1):
# Slice out appropriate part of array input_data by
# defining start:end in both dimensions
istart = istart
iend = istart + n1_new - 1
jstart = 0
jend = input_data.shape[0]
filename = "file_new_" + str(j) # output filename
# Write data to file
np.save( filename, input_data[jstart:jend, istart:iend] )
# Update starting point in input_data array
istart = istart + n1_new
# end for
# Finish last file (if necessary)
if (iend < input_data.shape[1]):
j = nfiles_new - 1
filename = "file_new_" + str(j) # output filename
# Slice out appropriate part of array input_data by
# defining start:end in both dimensions
istart = iend + 1
iend = input_data.shape[1]-1
jstart = 0
jend = input_data.shape[0]
# Write data to new file
np.save( filename, input_data[jstart:jend, istart:iend] )
# end ifThis code is a little longer than that to generate the data, but it does capture the case where the data is not evenly distributed across partitions, leaving the last partition smaller than the others.
As a note, I know my Python style is not Pythonic and that many may take issue with my coding style. That is perfectly fine, but please remember that this is just a quick example to illustrate what could be done. If you see an error, please comment on Twitter to the account adminHPC.
This example is a single process for redistributing the data across files and assumes that the single process can accommodate all necessary data in memory. If that’s not possible, you will have to write the parallel data redistribution code.
You could even contemplate writing code that reads only a small number of partitions into memory before writing new data partitions. The approach depends on the number of existing files and the number of “new” files needed, but the logic is approachable.
You might have to consider writing an application that uses more than one process if the amount of data cannot be stored in memory all at once.
Subset of Computational Processes Doing I/O
The other option for situations in which you have a different number of processes than files, preventing you from running one file per process, is to modify the computational code.
The computational code would have to read Mfiles files when it is using Nproc processes. The code would have to account for the case of Nproc > Mfiles and Mfiles > Nproc. To simplify this article, I will only consider the Nproc > Mfiles case.
The Nproc processes would have to read the Mfiles data files, determine which process needs what data, and send that data to the appropriate process (point-to-point communication). Some of these processes might be performing I/O themselves, so coordinating send and receive would be paramount to ge correct results, never mind performance.
Once all of the nodes have their data, computations can proceed. The final I/O can either be one file per process, as discussed previously, or a subset of Nproc.
Alternative: Single File for I/O
A degenerate case for the two previous paradigms, redistributing data with a separate application and using a subset of the computational processes for I/O, would be when Mfiles = 1. That is, a single file. I have seen and worked with code that runs with Nproc processes, but all I/O is performed by the rank 0 process (one file, or Mfiles = 1).
Obviously, this removes any benefits of parallel I/O because all I/O occurs on a single file. However, it does allow the application to scale with computational resources if the I/O is not a dominant part of total time. Remember, eventually Amdahl’s Law will get you, and the application will get less and less benefit from more processes until adding resources slows down the application.
The applications I’ve worked with rely on the rank 0 process having enough memory so it can read the input data before sending it to the other processes and so all the data can be received before writing. This solution works great when you have enough memory, but it’s not so easy when you don’t.
If you don’t have enough memory, you would have to develop a scheme wherein, for reading, the rank 0 process only reads a portion of the input data and sends it to the other processes. Then it would have to continue in this way until all of the data is read. When writing, the non-rank 0 processes only send a portion of their data to the rank 0 process, which then writes it to the file. This procedure is repeated until the data from all nodes is written. You would have to work out this scheme for your application.
MPI-IO
The MPI protocol is the de facto standard for writing distributed applications in HPC. It is also used in deep learning applications of which Horovod is an example. Initially, MPI was focused on sharing data and performing distributed computations, such as reductions, but over time, people wanted to use MPI for I/O, as well, and be able to write from distributed processes to a single shared file on a parallel filesystem.
In MPI-2, MPI-IO was adopted, adding functions that abstract I/O and allowing files to be accessed in a “patterned” way -- something that derived types did not facilitate. Note that MPI-4 is the current standard, of which MPI-IO is a part. With the use of collective I/O, MPI-IO has become a very powerful abstraction for improving I/O operations of distributed applications.
Classically, all processes (all ranks in the MPI communicator) participate in the I/O, but it’s not required. An application developer can choose to have a subset of a process perform I/O on behalf of all of the processes. You can define a new communicator, of which the processes perform I/O, or you can define multiple communicators to write to different files at the same time.
MPI-IO has become powerful enough that it now underlies high-level libraries (e.g., HDF5 and Parallel-NetCDF), because of the collective I/O capabilities that use large and contiguous I/O operations, resulting in increased I/O bandwidth and reduced locking.
In this article, I won’t cover MPI-IO, but you can find many tutorials and examples online.
Higher Level I/O Libraries
Rather than code their own special I/O routines, people have been turning to the use of high-level I/O libraries, which has some advantages, beginning with the libraries being highly tuned for performance. The libraries also have created some abstractions so that you don’t have to do low-level coding for I/O. Moreover, these libraries are also widely use so other applications (e.g., visualization tools) can read your library.
In this section, I present two such high-level libraries: HDF5, and netCDF.
HDF5
During the early years of HPC, it felt like thousands of data formats were in use. As you would expect, these formats were incompatible with one other for the most part, so users started demanding compatible formats. The multitude of formats started coalescing to just a few formats. In the early 1990s, NASA selected the Hierarchical Data Format (HDF) as the standard.
HDF5 is now one of the most common data formats for scientific and engineering applications. The current version of the HDF format is HDF5, and the most recent version is v5-1.12.2 released April 19, 2022. Rather than write your own code to read or write HDF5 files, you can link your code to libraries with an API for reading, writing, or manipulating data.
As a “universal” data format, HDF5 is self-describing. In essence, the data file can describe itself to the application by allowing an application to open the file with no a priori knowledge of the data or layout of the file and gather information on both the structure and content. Your application will use an API call to interrogate the file to understand its layout and metadata, including data labels.
For storing data, HDF5 uses the concept of objects, including (1) datasets, which are typed multidimensional arrays, and (2) groups, which are container structures that can hold datasets and other groups. With these two objects, HDF5 can create “… a truly hierarchical, filesystem-like data format.”
To use HDF5 effectively, you need to know something about the format, for which you can find tutorials and examples online. HDF5 is supported by many languages (e.g., C, Fortran, Julia, Go, MATLAB, Mathematica, Python, Perl, R, Rust, and Java). It also has tools to help manage the data files.
HDF5 is very popular in the scientific and engineering world and is widely used; however, it is also probably the most common data format for deep learning (DL) applications such as TensorFlow and PyTorch.
In general, you use HDF5 in your applications by having one of the N processes open the file and perform the I/O; therefore, that one process must be able to gather the data from the other processes and write it to the file. It’s not really performing parallel I/O but HDF5 has a parallel capability.
Parallel HDF5
The HDF5 parallel capability is named, aptly enough, Parallel HDF5. It is compatible with MPI and uses MPI-IO, allowing MPI applications to perform I/O to a single file, regardless of the number of processes. An application that uses Parallel HDF5 opens a shared file with an MPI communicator. All the processes in that communicator must participate in collective Parallel HDF5 APIs. If you want to open different files, then you can define a different communicator.
By using Parallel HDF5, you can use parallel I/O to achieve large speedups in I/O.
netCDF
Another popular data format is netCDF, which grew out of the atmospheric sciences arena, principally from University Corporation for Atmospheric Research (UCAR), and supports array-oriented scientific data. As with HDF5, it is a self-describing format. It provides interface libraries (APIs) for C, C++, Fortran77, and Fortran90, along with interfaces for R, Perl, Python, Ruby, Haskell, Mathematica, MATLAB, Julia, Octave, and Java.
netCDF has its own file format, generically described as the Common Data Format (CDF), but in version 4.0, netCDF added netCDF/HDF5, with some exceptions to the HDF5 file format.
All of the formats have a header that describes the layout of the file (data arrays and metadata). The files are platform independent with endianness addressed by the libraries.
As with HDF5, netCDF has some tools that can be used for file management, manipulation, and inspection. These tools are very useful in understanding what is contained in files and even visualizing the data.
You would use netCDF as you do HDF5. A single process within the N processes reads the data in the file and sends it to the other processes. When writing to a file in the netCDF format, the one process receives data from the other processes and writes it. It’s not parallel, but netCDF does have parallel capabilities.
Parallel-NetCDF
Around 2001, work began on a parallel implementation of Parallel-NetCDF that uses MPI-IO underneath. The initial goal was to focus on older formats: CDF-1, the “classic” format primarily used for netCDF through version 3.5, and CDF-2, added in CDF-1 version 3.6.0 onward, which allows even 32-bit applications to write files larger than 2GiB. In Parallel-NetCDF, only classic and 64-bit offset formats can be addressed in the CDF format.
Beginning with netCFD-4.0, Parallel-NetCDF can perform parallel I/O to HDF5 files. From version 4.1.1 onward, it supports parallel I/O to classic and 64-bit offset files with the Parallel-NetCDF library, but also the netCDF native APIs (i.e., you don’t have to use Parallel-NetCDF).
Parallel-NetCDF relies on an underlying MPI-IO library for performing the actual I/O.
Parallel Filesystems
Up to this point in the article I haven’t talked about filesystems. However, when I/O becomes parallel, the filesystem has a large effect on performance. HDF5, netCDF, and their parallel versions can be used on POSIX filesystems on Linux and the files will be correct. Performance is a different question.
Primarily, a performance difference arises between parallel filesystems such as Lustre or BeeGFS, network filesystems (i.e., NFS), or local filesystems. To help explain some of the issues, a very good paper by Rob Latham, Rob Ross, and Rajeev Thakur came out of Argonne National Labs titled “The Impact of File Systems on MPI-IO Scalability.” The paper is from 2004 but it nonetheless presents some interesting considerations, including:
The NFS consistency semantics work well in the serial environment for which they were designed, but they are a poor fit for parallel I/O. (pg. 3)
In essence, don’t expect great performance from NFS but all I/O will run correctly.
The authors also listed a few filesystem features that will help parallel I/O performance with MPI-IO. From the paper:
We have pointed out some characteristics of file system APIs and semantics that more effectively serve as the basis for MPI-IO implementations: support for noncontinuous I/O, better consistency semantics, client-independent file references, a stateless I/O model, and a caching model that allows a single file system operation to sync to storage … . By building parallel file systems with these characteristics in mind, MPI-IO implementations can leverage MPI collective communication and achieve good performance from the parallel file system even as the number of clients increases. (pg. 9)
Note that PVFS2 was the filesystem they used for testing ideas to improve MPI-IO performance.
Summary
Home systems now come with six or more cores. Clusters number thousands of nodes. These resources beg you to take advantage of them. However, staring you in the face, is Gene Amdahl telling you that the serial portion of an application limits how far it can be scaled (how many processors can be used). He’s not being mean; he’s trying to help you.
Applications have lots of sources of serial computation. One part that can cause difficulty is I/O. I’ve talked a bit about ideas for reducing the I/O time or having I/O time scale with the number of processes. One of the simplest ways is to have each process read and write their own data file. This approach is easy to code and scales with the number of processes, but it does limit you to only running on the same number of processes the next time and can cause issues with postprocessing applications that can’t read multiple files. People will disagree even mentioning this option for parallel I/O, but if they are so dead set against it, perhaps they would be willing to add the logic to your code for parallel I/O. By the way, you can write some tools to handle aggregating the files or partitioning them into a different number.
Another option is to use MPI-IO, which allows all processes in a MPI communicator to participate in the application I/O. This choice is a great way to achieve some level of parallel I/O, but the coding isn’t the easiest.
A good way to make your data files portable and reduce the I/O burden on yourself, is to use a common data format tool such as HDF5 or netCDF. The libraries and tools are self-describing making them very portable, and many postprocessing tools understand these types of files. Even better, you can use the parallel versions Parallel HDF5 and Parallel-NetCDF. These options use MPI-IO underneath the libraries to achieve parallel I/O and allow your application to scale better.