« Previous 1 2 3 Next »
pyamgx – Accelerated Python Library
Evolution of Scientific Languages
Scientific applications and languages have evolved over time. At first code was written and used by code developers, scientists, and engineers. They knew the code inside out because they wrote it, which also meant they could modify it when needed. At the same time, it meant they developed code instead of using the code to solve problems.
These initial programs were written in compiled languages such as Fortran and, later, C. The developers need to know about the underlying operating system and compilers. With this knowledge, they could tune the code for the best possible performance.
After awhile, code became more standardized, as did operating systems. Compilers were improved and could optimize for the more standardized code and operating systems. The same compiled languages, Fortran and C, were still being used.
At this point, people divided into developers and users. The developers mainly wrote and tested the applications, and users primarily focused on exercising the applications to solve problems and answer questions. Although there was still a fair amount of crossover between the groups, you could see each group focus on its primary mission.
The development and use of applications continued to evolve. Universities, for the most part, stopped teaching the classic compiled languages, except in computer science or specialized classes, and started teaching interpreted languages. These languages were easier to use and allowed users to develop applications quickly and simply. The thinking was that the syntax for these languages was simpler than compiled languages, and it was easier to test portions of the code because they were portable across operating systems and did not require knowledge of the compiler switches to create an executable. Many of these interpreted languages also had easy-to-use “add-ons,” or libraries, that could extend the application to use a GUI, plot results, or provide extra features without writing new code.
By using these interpreted languages, the user could focus more on using the application because coding was easier. Of course, the goal of using the applications was to get the answers as fast and as accurately as possible. The use of interpreted languages allowed the users either to assemble their applications from libraries quickly or just to use prebuilt applications that were simple to understand or modify. Overall, the applications might run more slowly than a compiled application, but the total time to start running an application was less (i.e., less time to science).
With the use of languages such as Python, Julia, or Matlab, applications can be created quickly. Moreover, the parts of the application that create bottlenecks can use accelerated libraries to reduce or even eliminate the bottlenecks.
A common approach to marrying libraries with interpreted languages is to create “wrappers,” so the libraries can be imported and used. This approach leaves the development of the underlying libraries without having to worry about interpreted languages.
A classic example of this approach is Python. A number of libraries – not just accelerated libraries – have been wrapped and are available to be “imported” into Python. Examples include:
- CuPy
- TensorFlow
- cuBLAS
- cuFFT
- Magma
- AWS
- Plotly
- OpenMesh
- Trimesh (loading and using triangular meshes)
- MeshPy (meshing for partial differential equations)
- scikit-cuda
- PyIMSL
pyamgx Interface
Code developers have gotten used to developing Python interfaces to libraries, but some libraries do not have a Python interface. AmgX was one example, so Ashwin Srinath at Clemson University wrote an interface. Called pyamgx, it allows you to use AmgX in Python applications using some very simple function calls (i.e., it is very Pythonic).
From the documentation, a simple example would be the following (the SciPy version of the code is compared with the AmgX results):
# Create matrices and vectors:
A = pyamgx.Matrix().create(rsc)
x = pyamgx.Vector().create(rsc)
b = pyamgx.Vector().create(rsc)
# Create solver:
solver = pyamgx.Solver().create(rsc, cfg)
# Upload system:
M = sparse.csr_matrix(np.random.rand(5, 5))
rhs = np.random.rand(5)
sol = np.zeros(5, dtype=np.float64)
A.upload_CSR(M)
b.upload(rhs)
x.upload(sol)
# Setup and solve:
solver.setup(A)
solver.solve(b, x)
# Download solution
x.download(sol)
print("pyamgx solution: ", sol)
print("scipy solution: ", splinalg.spsolve(M, rhs))To help people understand how they can take advantage of pyamgx, Ashwin created a Jupyter notebook that uses pyamgx with FiPy, a finite volume PDE solver built in Python.
The notebook goes over an example of solving the diffusion equation for simple 2D problems. FiPy can use solvers from SciPy, which is used for the CPU example of a steady-state problem. The solver uses the generalized minimal residual method (GMRES) solver from the pyamgx package (it is the default solver). The notebook also uses the conjugate gradient squared (CGS) solver to illustrate how different solvers can be used.
The next section of the notebook discusses how to use pyamgx within FiPy by defining a custom PyAMGXSolver class; then, the notebook runs FiPy with AmgX over a range of grid sizes. It also solves the problem using the CPU solver from SciPy. Finally, a comparison of the compute time for the CPU and the GPU is made and plotted.
Features Ashwin added to pyamgx are routines that allow NumPy arrays and SciPy sparse matrices that reside in CPU memory to be “uploaded” to the GPU. On the GPU, these are AmgX vectors and matrices. You can also “download” the AmgX vectors that are on the GPU back to NumPy arrays that are on the CPU.
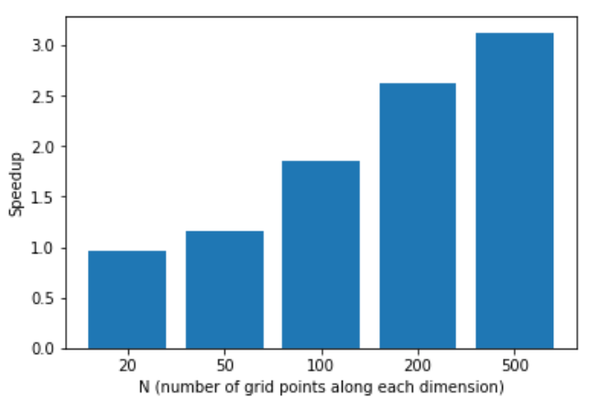
In case you are curious, at the end of Ashwin’s notebook, he plots the speedup using AmgX. The GPU was an Nvidia PCIe P100 (I do not know what CPUs were used). Figure 1 shows the speedup as a function of the problem size dimension (it was a square grid).
When the problem dimension reaches 500 (i.e., 500x500, or 250,000 grid points), the pyamgx wrapper around AmgX speedup is 3x.
« Previous 1 2 3 Next »